Analysis of activator and repressor functions reveals the requirements for transcriptional control by LuxR, the master regulator of quorum sensing in Vibrio harveyi
Abstract
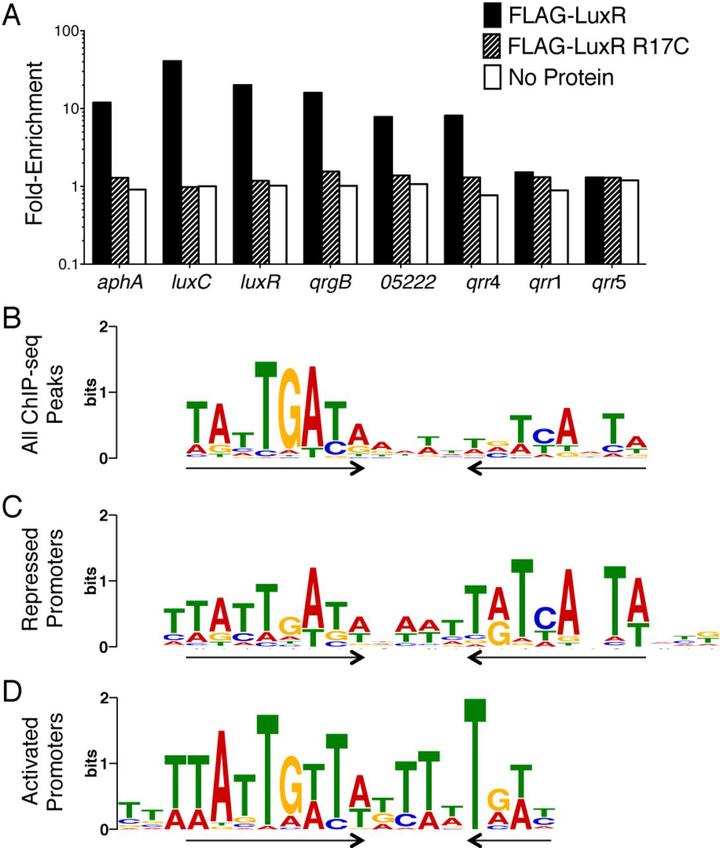
LuxR-type transcription factors are the master regulators of quorum sensing in vibrios. LuxR proteins are unique members of the TetR superfamily of transcription factors because they activate and repress large regulons of genes. Here, we used chromatin immunoprecipitation and nucleotide sequencing (ChIP-seq) to identify LuxR binding sites in the Vibrio harveyi genome. Bioinformatics analyses showed that the LuxR consensus binding site at repressed promoters is a symmetric palindrome, whereas at activated promoters it is asymmetric and contains only half of the palindrome. Using a genetic screen, we isolated LuxR mutants that separated activation and repression functions at representative promoters. These LuxR mutants exhibit sequence-specific DNA binding defects that restrict activation or repression activity to subsets of target promoters. Altering the LuxR DNA binding site sequence to one more closely resembling the ideal LuxR consensus motif can restore in vivo function to a LuxR mutant. This study provides a mechanistic understanding of how a single protein can recognize a variety of binding sites to differentially regulate gene expression.